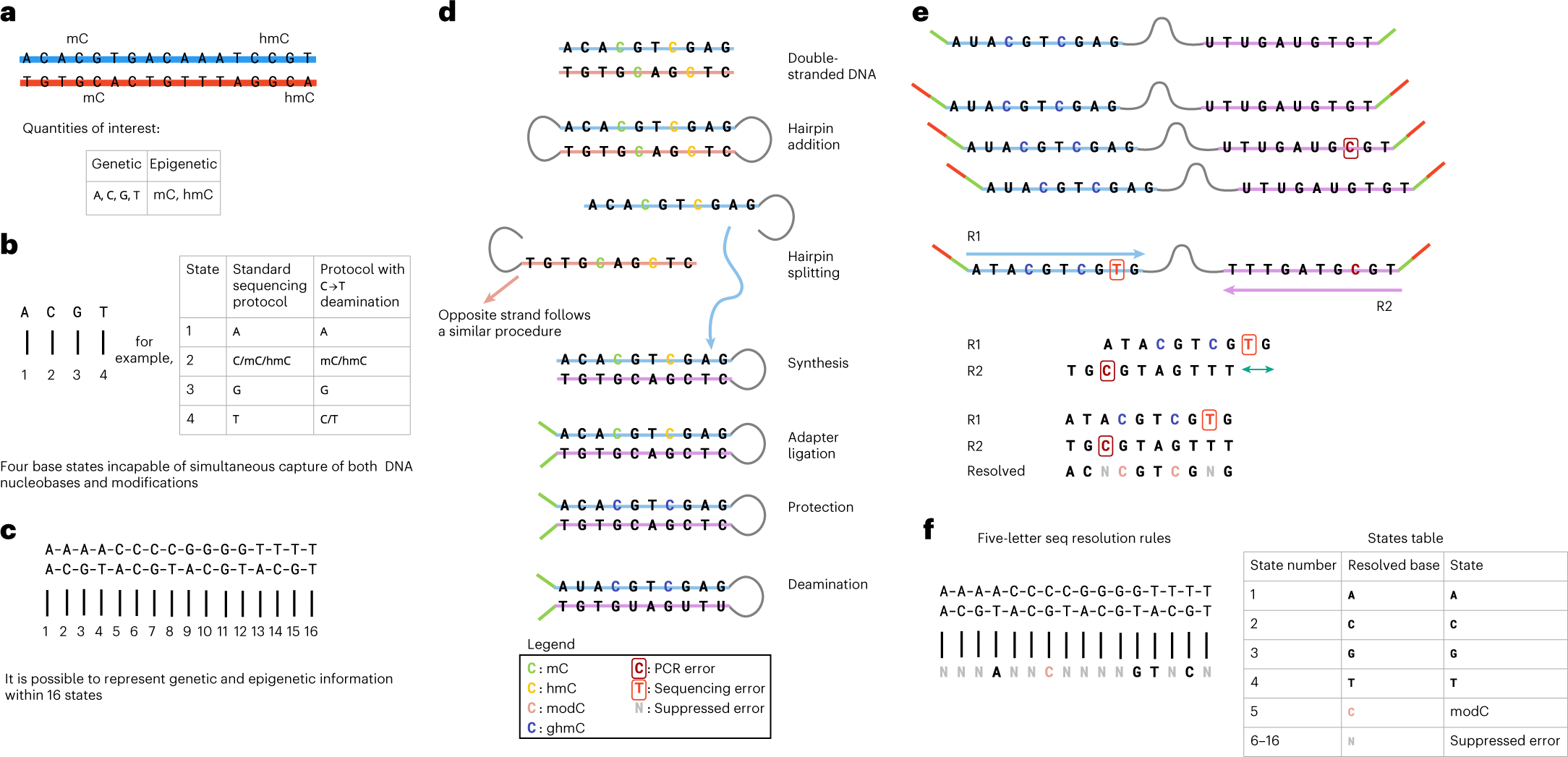
Quantification of epigenetic biomarkers: an evaluation of established and emerging methods for DNA methylation analysis | BMC Genomics | Full Text
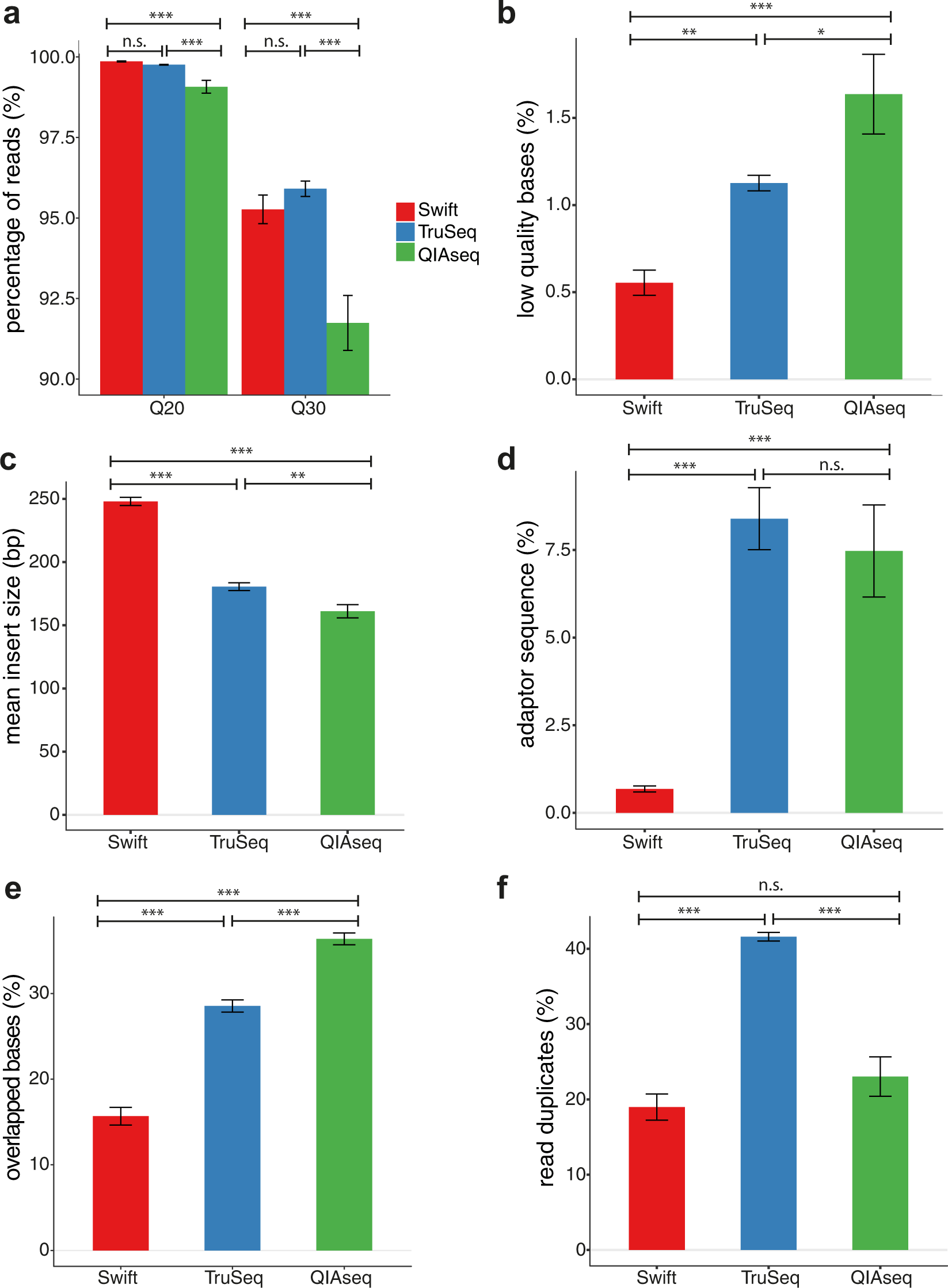
Systematic evaluation of library preparation methods and sequencing platforms for high-throughput whole genome bisulfite sequencing | Scientific Reports

Comparison of whole-genome bisulfite sequencing library preparation strategies identifies sources of biases affecting DNA methylation data | bioRxiv
Circulating cell-free DNA from plasma undergoes less fragmentation during bisulfite treatment than genomic DNA due to low molecular weight | PLOS ONE

Base-resolution profiling of active DNA demethylation using MAB-seq and caMAB-seq | Nature Protocols
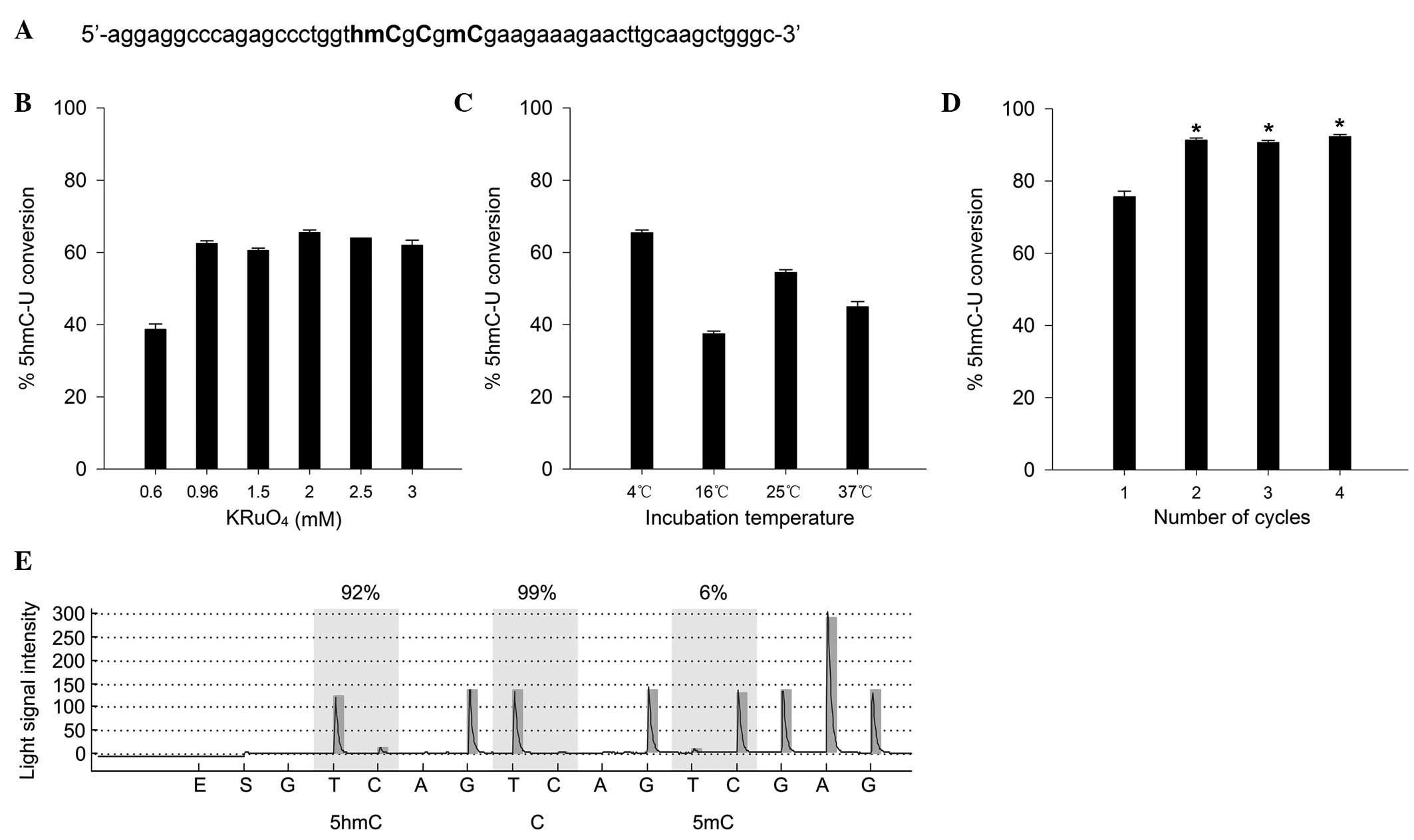
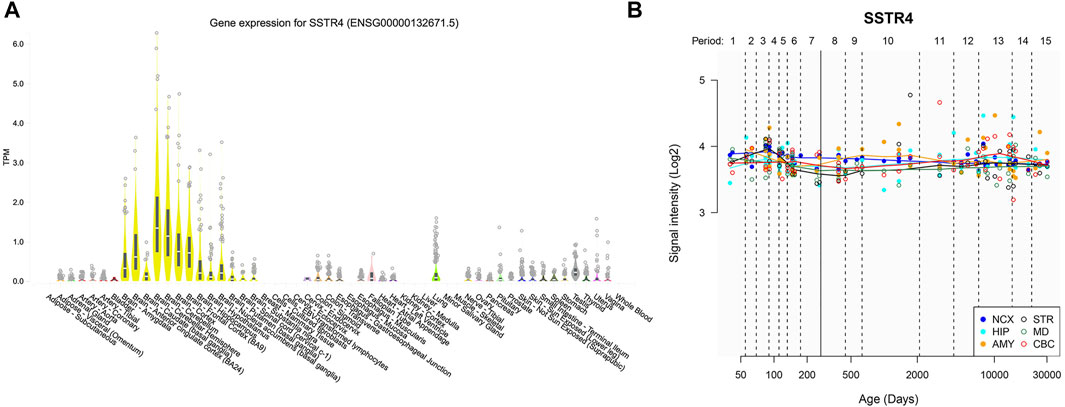
PRDM8 internal promoter hyperhydroxymethylation correlates with increased expression of the corresponding transcript in Down syndrome
Global DNA methylation and chondrogenesis of rat limb buds in a three-dimensional organ culture system | Biomolecules and Biomedicine
Full article: Comparative analysis of 12 different kits for bisulfite conversion of circulating cell-free DNA
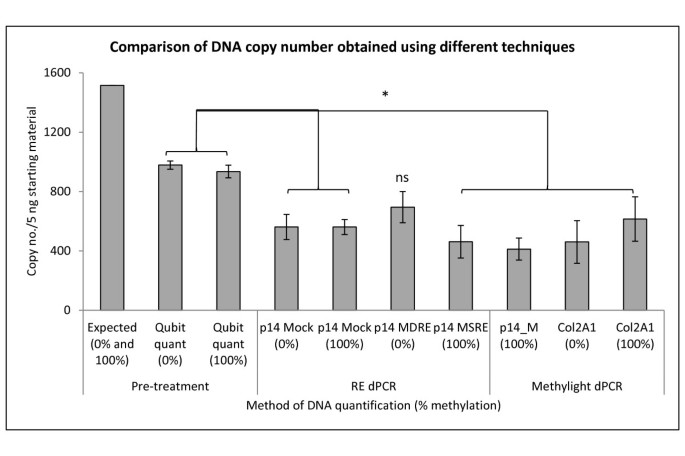
Evaluation of bisulfite kits for DNA methylation profiling in terms of DNA fragmentation and DNA recovery using digital PCR | PLOS ONE
Table 1: Protocol steps and requirements for CEF workflow Starting material One to two 5 µm thick sections from tumor tissue FF