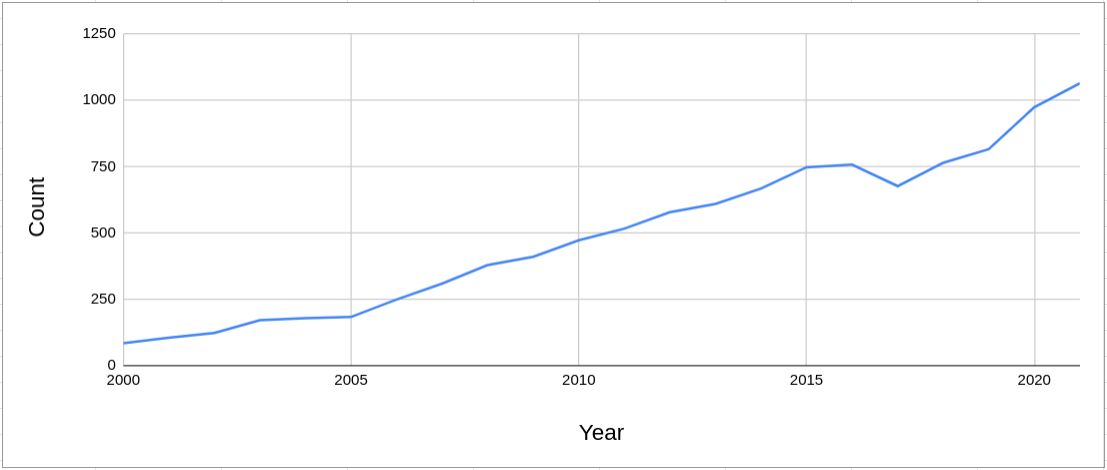
ploidy and copy number states very different between SNParray (ASCAT) and WGS (FACETS) · Issue #113 · mskcc/facets · GitHub
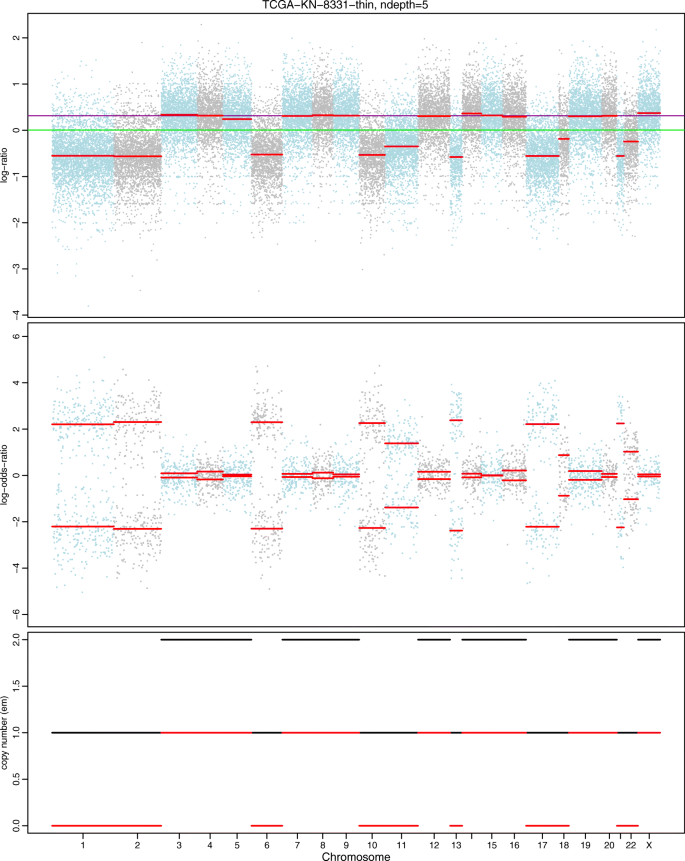
hsegHMM: hidden Markov model-based allele-specific copy number alteration analysis accounting for hypersegmentation | BMC Bioinformatics | Full Text

Copy number estimates from FACETS for Patient 1. A decrease in copy... | Download Scientific Diagram
![PDF] FACETS: allele-specific copy number and clonal heterogeneity analysis tool for high-throughput DNA sequencing | Semantic Scholar PDF] FACETS: allele-specific copy number and clonal heterogeneity analysis tool for high-throughput DNA sequencing | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/0333c81df745873f6cadadb6009446166eea7136/5-Figure4-1.png)
PDF] FACETS: allele-specific copy number and clonal heterogeneity analysis tool for high-throughput DNA sequencing | Semantic Scholar
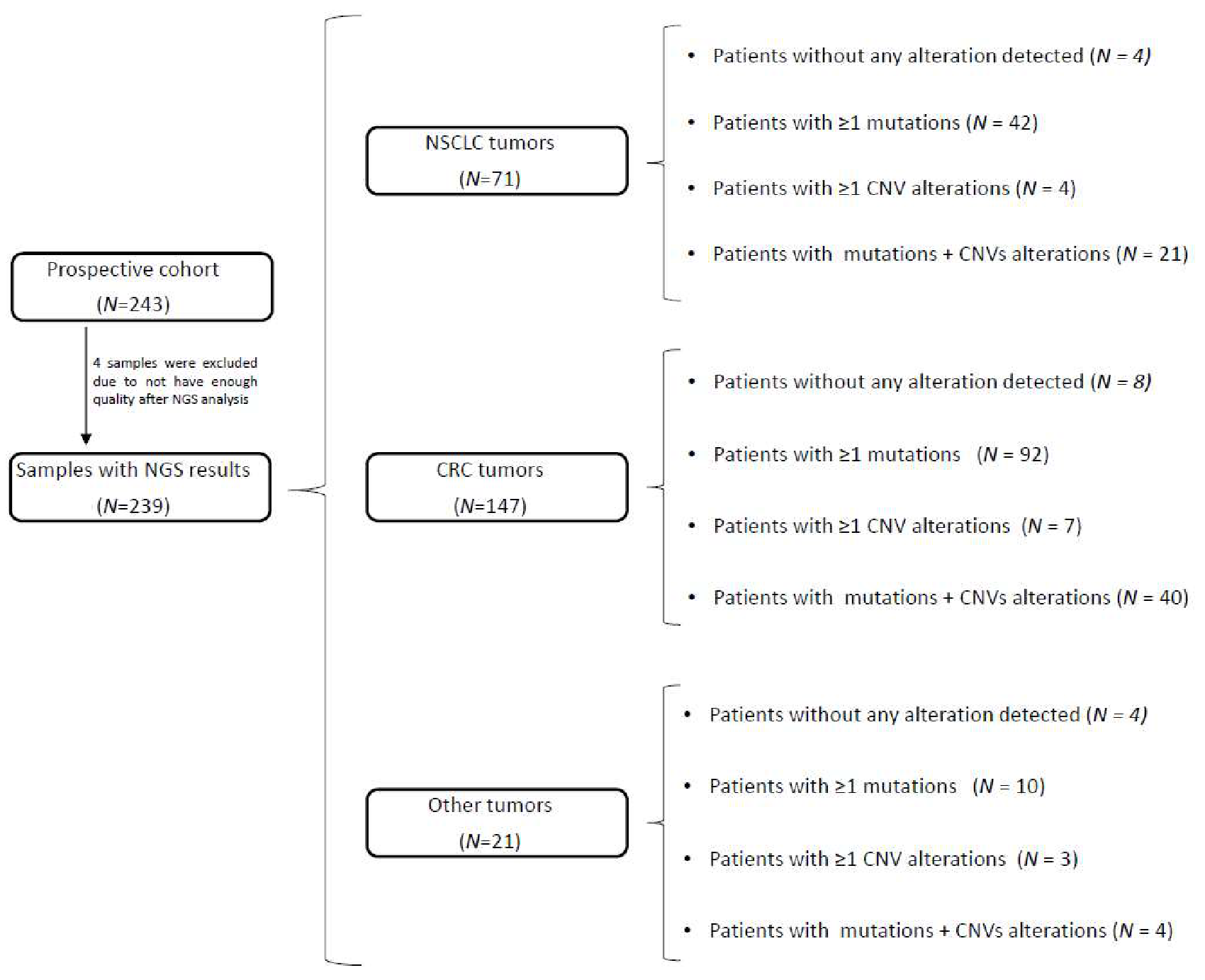
JMP | Free Full-Text | Analysis of Copy Number Variations in Solid Tumors Using a Next Generation Sequencing Custom Panel
Copy number signature analysis tool and its application in prostate cancer reveals distinct mutational processes and clinical outcomes | PLOS Genetics
![PDF] FACETS: allele-specific copy number and clonal heterogeneity analysis tool for high-throughput DNA sequencing | Semantic Scholar PDF] FACETS: allele-specific copy number and clonal heterogeneity analysis tool for high-throughput DNA sequencing | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/0333c81df745873f6cadadb6009446166eea7136/2-Figure1-1.png)
PDF] FACETS: allele-specific copy number and clonal heterogeneity analysis tool for high-throughput DNA sequencing | Semantic Scholar

Detection of Biallelic Loss of DNA Repair Genes in Formalin-Fixed, Paraffin-Embedded Tumor Samples Using a Novel Tumor-Only Sequencing Panel - The Journal of Molecular Diagnostics
Diversity of Prdm9 Zinc Finger Array in Wild Mice Unravels New Facets of the Evolutionary Turnover of this Coding Minisatellite | PLOS ONE

Copy number signatures predict chromothripsis and clinical outcomes in newly diagnosed multiple myeloma | Nature Communications

Figure 7 from FACETS: allele-specific copy number and clonal heterogeneity analysis tool for high-throughput DNA sequencing | Semantic Scholar
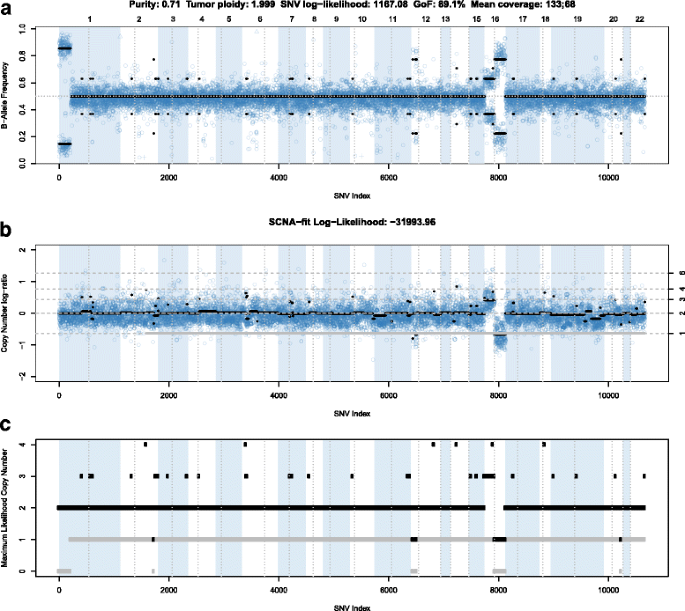
PureCN: copy number calling and SNV classification using targeted short read sequencing | Source Code for Biology and Medicine | Full Text

Global diversity, population stratification, and selection of human copy- number variation | Science

ploidy and copy number states very different between SNParray (ASCAT) and WGS (FACETS) · Issue #113 · mskcc/facets · GitHub

Comprehensive Assessment of Somatic Copy Number Variation Calling Using Next-Generation Sequencing Data | bioRxiv

Accurate quantification of copy-number aberrations and whole-genome duplications in multi-sample tumor sequencing data | Nature Communications